|
Issue 36
Transport Phenomena Archive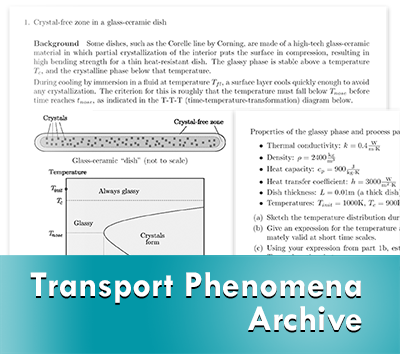
We are pleased to announce that the Transport Phenomena Archive has been updated and published on nanoHUB. It was originally built by Adam Powell and Matthew Krane in the mid-2000s as a resource facilitating the work of materials transport educators and students in formal and informal settings. The archive consists of high-quality Open Educational Resources (OERs), including readings, problems, videos, and open source codes.
The Archive was uploaded and published on nanoHUB by Dr. Powell in collaboration with Joe Cychosz, NCN's Production Manager, and supported by a Faculty Learning Community Teaching Innovation Grant that allowed for enhancement of the resources.
According to Dr. Powell, it is important to note that the OERs, "...are (almost all) copyrighted, and their licenses place some limitations on what you can do with them. You may freely read them, print and copy them, share them with friends, and use them in classes as you like. If you distribute them, be sure to include copyright information, and make sure you're abiding by license limitations – or contact the authors regarding separate licensing."
Adam Powell is an Associate Professor in the Mechanical Engineering department at Worcester Polytechnic Institute. His research interests include using materials processing, particularly the tools of electrochemistry and process modeling, to eliminate barriers to reducing, eliminating, and reversing greenhouse emissions.
Visit the Transport Phenomena Archive Award Winning Researchers
Congratulations to nanoHUB's own Gerhard Klimeck, Director of the Network for Computational Nanotechnology! Dr. Klimeck received the prestigious Humboldt Research Award on June 27th at Charlottenburg Palace in Berlin, Germany.
More information: https://www.purdue.edu/newsroom/...

We would also like to congratulate Paul Macklin, associate professor of intelligent systems engineering and nanoBIO node researcher, who won the 2019 Research Prize in the category of public impact from the journal Public Library of Science (PLoS) Computational Biology.
Dr. Macklin's PhysiCell framework, which allows researchers to model how cells, for example, may react to the introduction of drugs to combat cancer or other diseases, is used widely in nanoHUB simulation tools.
More information: https://sice.indiana.edu/news/...
PhysiCell-based tools on nanoHUB: https://nanohub.org/search/?terms=physicell Share Your Feedback on the nanoHUB Website
Would you like to help improve nanoHUB? We're planning to conduct focus groups regarding the usability of the website in the near future. If you'd like to take part, please contact:
Brad Hallberg
Business Development Specialist
hallberg@purdue.edu
Upcoming Events
Micro Nano Technology Education Special Interest Group - Face-to-Face Meeting at High Impact Technology Conference
When:
Tuesday, July 23, 2019
9:30 a.m. - 5:30 p.m. EST
Where: St. Louis, Missouri
The MNTeSIG will be held July 23, 2019, during the High Impact Technology Exchange Conference (HI-TEC) in St. Louis, Missouri (July 22-25, 2019). The Micro Nano Technology education Special Interest Group is a collection of educators and industry partners who work together throughout the year to bring micro and nanotechnology to classrooms across the US and internationally.
Website: Micro Nano Technology Education Special Interest Group
NANOTECH 2019: A Collective Approach for Innovation and Invention
When:
Sunday, July 28 - Monday, July 29, 2019
Where: Paris, France
Nanotech 2019 will be a platform to integrate and motivate participants through presentations, exhibition, papers, workshops, a poster session, symposium, and panel discussion. Nanotechnology and nanomaterials are the latest trending technologies in many fields especially in industry sectors such as information technology, homeland security, medicine, transportation, energy, food safety, and environmental science.
Registration and abstract submission are now open.
Website: NANOTECH 2019

New Resources
This course is an introduction to the fundamentals of fiber optic communications, which constitute the backbone of the internet. The course will start with a refresher on the operation of key components needed for an effective fiber optic communication system, and then show how these components interact at a system level. Finally, the course will conclude with outlook for future research in extending the capabilities of these networks to higher bandwidths and quantum-secured communications.
The NSF nanomanufacturing (nanoMFG) node, hosted at the University of Illinois at Urbana-Champaign, presented its first workshop exploring the potential opportunities for an emerging area at the intersection of data science and nanomanufacturing. This workshop brought together experts in nanomanufacturing, data science, and cyberinfrastructure. The workshop centered on the opportunities to leverage the following three themes in nanomanufacturing:
- Data opportunities and challenges in nanomanufacturing processes, characterization, and metrology
- Data infrastructure: real-time collection, storage, and sharing
- Data-enabled modeling, large-scale computing, and machine intelligence
The Spin Qubit Lab calculates the device-level characteristics of spin-based quantum gates. Single-qubit rotational gates and two-qubit controlled-phase gate can be simulated, which form a complete set of quantum gates for universal quantum computing. The tool can simulate time evolution, compute delay and fidelity, and perform quantum process tomography for the quantum gates. The effect of dephasing can be simulated by inputting a spin dephasing time.
This application simulates coarse-grained ligand-receptor binding of ligand-decorated virus particles interacting with a cell wall. The viruses are modeled after P22 virus-like particles (VLPs) of diameter 60 nanometers, and ligands represent smaller nanoparticles of 6 nanometer diameter. Each virus is fully decorated with 60 ligands. Receptors are fixed on a lattice of tunable size.
This application was developed to simulate the liver tissue mechanobiology, and it successfully models the interaction between tumor cells and parenchyma cells in different biomechanical parameters. The software is powered by PhysiCell, a powerful simulation tool that combines multi-substrate diffusive transport and off-lattice cell models.
|






